MS/MS Mass Spectrometry Filtering Tree for Bile Acid Regio- and Stereoisomer Annotation
Anal. Chem. 2026, 98, 6, 4571–4584: Graphical abstract
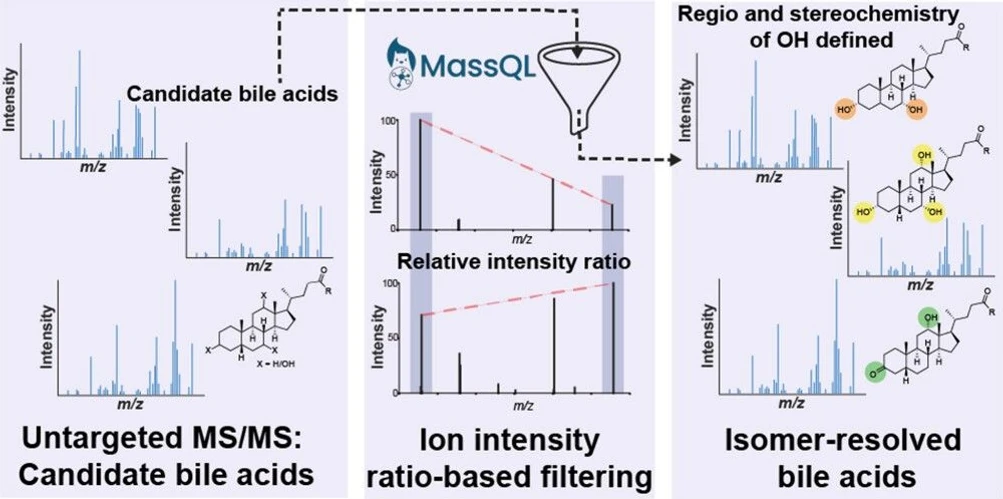
This study presents a novel MS/MS filtering tree approach for distinguishing regio- and stereoisomeric bile acids in untargeted LC-MS/MS data. By leveraging intensity ratios of closely related fragment ions, the method overcomes limitations of traditional spectral matching, enabling more accurate isomer annotation.
A user-friendly web-based application was developed to facilitate implementation without coding expertise. Applied to public datasets, the workflow revealed diet-associated bile acid patterns across mammalian species and enabled identification of previously uncharacterized compounds, demonstrating its potential for large-scale metabolomics studies.
The original article
MS/MS Mass Spectrometry Filtering Tree for Bile Acid Regio- and Stereoisomer Annotation
Ipsita Mohanty, Shipei Xing, Vanessa Castillo, Julius Agongo, Abubaker Patan, Yasin El Abiead, Helena Mannochio-Russo, Wilhan D. Gonçalves Nunes, Jasmine Zemlin, Itzhak Mizrahi, Dionicio Siegel, Mingxun Wang, Lee R. Hagey, and Pieter C. Dorrestein*
Anal. Chem. 2026, 98, 6, 4571–4584
https://doi.org/10.1021/acs.analchem.5c05677
licensed under CC-BY 4.0
Selected sections from the article follow. Formats and hyperlinks were adapted from the original.
Cholesterol-derived steroid molecules, known as bile acids, have been detected in nearly every organ where they have been studied. (1) Alterations in bile acid composition and dysbiosis have been linked to various health conditions, including neurocognitive developmental disorders, cancer, infections, and metabolic diseases such as diabetes and inflammatory bowel disease (IBD). (2−8) In the last 5 years, the number of detected bile acids in metabolomics experiments (1,9−15) has increased by the 1000s and possibly approaching 10,000s - most of which have not yet been fully structurally characterized. (1,6,16−19) We believe this is only the tip of the iceberg and that in the next 5 years this number will continue to increase. The scale of the bile acid pool and the challenge of accurately detecting and describing their structure truly represent an exciting frontier for analytical and computational scientists who continue to push the boundaries of what is possible.
In humans, bile acids are predominantly derived from cholesterol through pathways that exist not only in the liver as previously thought but also in many other organs including kidneys and brain. (16,20,21) These pathways shorten the side chain, producing a carboxyl terminus on the remaining 24-carbon steroid structure, which consists of four interconnected rings and side chain, collectively referred to here as the “core” (colored in Figure 1a). The core of bile acids undergoes multiple hydroxylations, a process typically associated with the liver but also occurring in other organs such as the brain, kidney, and spleen. (16) Considering hydroxylation at known carbon positions on bile acids reported from human samples (as starred in Figure 1a, Supplementary Table S1) along with stereoisomers and allo versus nonallo A/B ring isomerization at C5, approximately 1,800 candidate mono-, di-, and trihydroxy C24 bile acid cores are theoretically possible (Figure 1a). This number further expands exponentially in the gut.
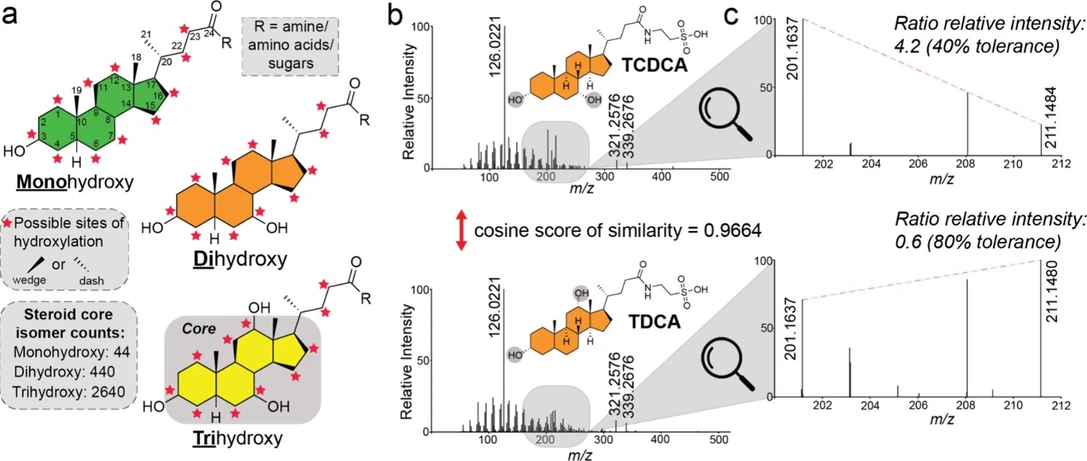 Anal. Chem. 2026, 98, 6, 4571–4584: Figure 1. MS/MS fragmentation of bile acid isomers. (a) Structure of bile acids highlighting mono-, di-, and trihydroxylated steroid cores, with experimentally observed potential hydroxylation sites on the steroid core indicated by red stars. (b) MS/MS fragmentation spectra of the regioisomers, taurochenodeoxycholic (TCDCA) acid and taurodeoxycholic acid (TDCA), illustrating a low-intensity mass region containing ions unique to each isomer. (c) Enlarged view of the MS/MS fragmentation spectra for taurochenodeoxycholic and taurodeoxycholic acids, emphasizing the ion pair used to calculate relative intensity ratios for differentiating these isomers.
Anal. Chem. 2026, 98, 6, 4571–4584: Figure 1. MS/MS fragmentation of bile acid isomers. (a) Structure of bile acids highlighting mono-, di-, and trihydroxylated steroid cores, with experimentally observed potential hydroxylation sites on the steroid core indicated by red stars. (b) MS/MS fragmentation spectra of the regioisomers, taurochenodeoxycholic (TCDCA) acid and taurodeoxycholic acid (TDCA), illustrating a low-intensity mass region containing ions unique to each isomer. (c) Enlarged view of the MS/MS fragmentation spectra for taurochenodeoxycholic and taurodeoxycholic acids, emphasizing the ion pair used to calculate relative intensity ratios for differentiating these isomers.
Recently, we developed MS/MS fragmentation-based filters using a Mass Spectrometry Query Language (MassQL), a query language that allows filtering of data patterns (49) to retrieve all MS/MS spectra from the Orbitrap data sets in GNPS/MassIVE repository containing bile acid-specific fragment ions. After merging similar spectra, this resulted in a reference library of 21,549 MS/MS spectra. (1) This library includes MS/MS spectra of known and many yet-to-be structurally characterized bile acid candidates that have already been detected from public metabolomics data repositories. This also includes MS/MS of multiple ion forms such as different adducts, in-source fragments, and multimers, (1) which all help in the annotation of bile acids. This MassQL library enables the annotation of the number of hydroxyl groups on the bile acid core, as well as the identification of potential modifications, and ∼ 62% of the atomic compositions are annotated. Other spectral “modifications” that do not have a distinct atomic composition may be different adducts or multiple combinations of adducts, in source fragments, and multimers. However, the diagnostic bile acid MS/MS fragments used to build this reference library do not allow for the differentiation between regio- and stereoisomers. Also, MS/MS spectral matching with the candidate library using the cosine metric often overlooks finer details within fragment ions, as minor peak intensity changes or low-intensity fragment ions have a minimal influence on overall spectral match scores (Figure 1b).
Based on our manual inspection of hundreds of bile acid MS/MS spectra, we observed and hypothesized that information in the low-intensity MS/MS fragments, including specific ion pairs, their intensity ratios, and unique ions, could be used to distinguish the steroid core isomers. As proof of concept in leveraging MS/MS data to enable further refinement of the stereo and regiochemistry of the hydroxylations of the bile acids cores, we developed a MassQL-based filtering tree utilizing key marker ions and their intensity ratios within narrow m/z windows to propose regio- and stereoisomer assignments of bile acids solely from MS/MS data. In this study, we focus on bile acid amidates, a class of structures found in humans and other animals.
Experimental Section
MS/MS Data Acquisition of Taurine-Conjugated Bile Acids
We generated one mM stock solutions of our in-house collection of 48 taurine-conjugated bile acids in 100% MeOH. The standards used in this study were obtained from the bile acid collection of the late Dr. Alan Hofmann’s laboratory at UCSD. The Dorrestein laboratory inherited these standards after his demise. The standard samples were injected (5 μL) into a Vanquish ultrahigh-performance liquid chromatography (UHPLC) system coupled to a Q-Exactive quadrupole Orbitrap mass spectrometer (Thermo Fisher Scientific, Waltham, MA). A Kinetex polar C18 column (2.1 × 100 mm, 2.6 μm particle size, 100 A pore size; Phenomenex, Torrance) was employed with a SecurityGuard C18 column (2.1 mm ID) at 40 °C column temperature. The mobile phases (0.5 mL/min) were 0.1% formic acid in both water (A) and ACN (B) with the following gradient: 0–0.5 min 5%B, 0.5–1.1 min 5–25%B, 1.1–7.5 min 25–40%B, 7.5–8.5 min 40–99%B, 8.5–10 min 99%B, 10–12 min 5%B. The mass spectrometer was operated in positive heated electrospray ionization with the following parameters: sheath gas flow, 53 AU; auxiliary gas flow, 14 AU; sweep gas flow, 3 AU; auxiliary gas temperature, 400 °C; spray voltage, 3.5 kV; inlet capillary temperature, 269 °C; S-lens level, 50 V. MS1 scan was performed at m/z 150–1500 with the following parameters: resolution, 35,000 at m/z 200; maximum ion injection time, 100 ms; automatic gain control (AGC) target, 1E6. Up to 5 MS/MS spectra per MS1 scan were recorded under the data-dependent mode (dd-MS2) with the following parameters: resolution, 17,500 at m/z 200; maximum ion injection time, 150 ms; AGC target, 5.0E5; MS/MS precursor isolation window, m/z 1; isolation offset, m/z 0.5; normalized collision energy (NCE) of 45%; minimum AGC for MS/MS spectrum, 2.5E4; apex trigger, 2 to 5 s; dynamic precursor exclusion, 8 s. Fragmentation was performed in the HCD cell using nitrogen as the collision gas. The data was deposited in GNPS/MassIVE and is publicly available at MSV000092003. The MS/MS spectra are also added now to the GNPS spectral library - BILELIB19 (https://gnps.ucsd.edu/ProteoSAFe/gnpslibrary.jsp?library=BILELIB19)
Results and Discussion
Retrospective Bile Acid Isomer Profiling in a Public Data Set of Animals with Different Diets
To illustrate the utility of the MassQL multistep filtering trees in retrospectively analyzing data in public repositories, we selected a publicly available data set containing feces from 40 animals (13 mammalian species) from the Zoological Center Tel Aviv-Ramat Gan, Israel. (62) The data set consisted of 3 herbivores, 23 omnivores, and 13 carnivores (Figure 4a). We selected this data set as we had previously confirmed detection of 6 previously uncharacterized polyamine-conjugated bile acids that we could not confirm with retention times as none of the standards we had synthesized matched, although their MS/MS did. We showed that their levels were higher in carnivores compared to herbivores and omnivores; (1) however, we were still unable to resolve the regio- and stereochemistry of all possible isomers of polyamine-conjugated bile acids. To further dive into the diversity of bile acids, we matched the MS/MS spectra of this study against the GNPS spectral libraries, including the candidate bile acid MS/MS reference library that we had previously created from the repository scale analysis. (1) This revealed 993 MS/MS spectral matches to bile acids. We next applied the multistep MassQL queries to resolve the regio- and stereochemistry of the bile acids. The steps described in this section follow the same filtering approach as outlined for the dihydroxy bile acid filtering tree (Figure 2).
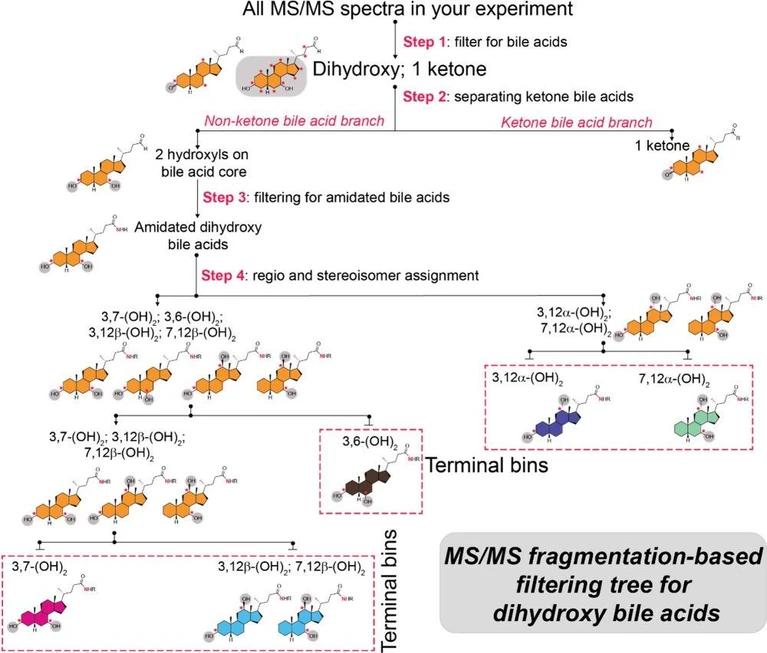 Anal. Chem. 2026, 98, 6, 4571–4584: Figure 2. Development of MS/MS fragmentation-based MassQL filtering tree. Sequential MS/MS fragmentation-based filtering tree designed to classify regio- and stereoisomers of dihydroxylated bile acids. Structures at each filtering step are shown, with terminal bins color-coded for clarity.
Anal. Chem. 2026, 98, 6, 4571–4584: Figure 2. Development of MS/MS fragmentation-based MassQL filtering tree. Sequential MS/MS fragmentation-based filtering tree designed to classify regio- and stereoisomers of dihydroxylated bile acids. Structures at each filtering step are shown, with terminal bins color-coded for clarity.
Step 1 excludes MS/MS spectral matches to candidate bile acids where the experimental spectra lacked the two diagnostic MS/MS ions for bile acids. Steroids and lipids often exhibit similar MS/MS spectra to bile acids, sometimes yielding high cosine similarity scores (>0.7), yet they lack key low-intensity diagnostic ions. Due to the low intensity of these ions, their presence or absence typically does not significantly affect spectral matching scores. However, our MassQL isomer filters in Step 1 identified these MS/MS spectra to reveal that 543 MS/MS spectra had the two characteristic fragment ions for mono-, di-, and trihydroxy bile acids (Figure 4a). A portion of the 543 MS/MS spectra are adducts, in-source fragments, and multimers, overall reducing the number of unique bile acid structures. Based on retention time and peak shape correlation analysis in MZmine4, (63) a strategy that allows the discovery of ion forms associated with a specific molecule, we estimate these to represent 402 bile acids. Even though the data contained evidence for hundreds of unique bile acids, the stereo- and regiochemistry of hydroxylations were yet to be defined.
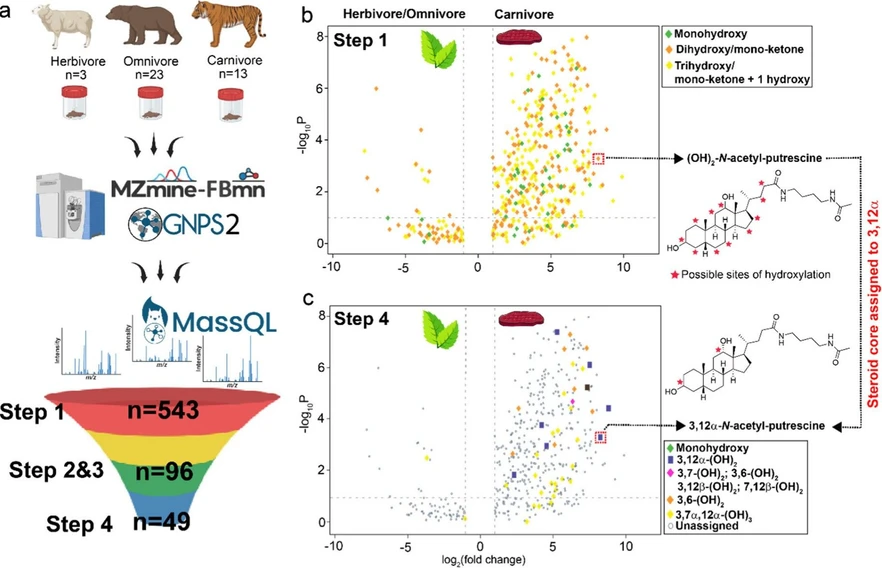 Anal. Chem. 2026, 98, 6, 4571–4584: Figure 4. Filtering of bile acid isomers in an untargeted LC-MS/MS data set. Representative example for using the MassQL filters for identifying bile acid isomers. (a) Overview of study design. Fecal samples from herbivores (n = 3), omnivores (n = 23), and carnivores (n = 13) were analyzed by LC-MS/MS and processed using MZmine4 and FBMN on GNPS2. Stepwise MassQL filtering refined candidate bile acids from 543 features (Step 1) to 96 (Steps 2 and 3) and 49 stereo and regiochemistry assigned MS/MS spectra (Step 4). Differential abundance of bile acids between herbivores/omnivores and carnivores is illustrated in the volcano plots (b) at Step 1 and (c) Step 4 of the dihydroxy filtering tree. The annotation of a previously uncharacterized bile acid was refined from (OH)2-N-acetyl-putrescine to 3,12α-N-acetyl-putrescine.
Anal. Chem. 2026, 98, 6, 4571–4584: Figure 4. Filtering of bile acid isomers in an untargeted LC-MS/MS data set. Representative example for using the MassQL filters for identifying bile acid isomers. (a) Overview of study design. Fecal samples from herbivores (n = 3), omnivores (n = 23), and carnivores (n = 13) were analyzed by LC-MS/MS and processed using MZmine4 and FBMN on GNPS2. Stepwise MassQL filtering refined candidate bile acids from 543 features (Step 1) to 96 (Steps 2 and 3) and 49 stereo and regiochemistry assigned MS/MS spectra (Step 4). Differential abundance of bile acids between herbivores/omnivores and carnivores is illustrated in the volcano plots (b) at Step 1 and (c) Step 4 of the dihydroxy filtering tree. The annotation of a previously uncharacterized bile acid was refined from (OH)2-N-acetyl-putrescine to 3,12α-N-acetyl-putrescine.
The application of our MassQL filtering tree allowed us to further refine structural details within the bile acid core. After Step 2 and Step 3 of the MassQL filtering, we separated the amidated mono-, di-, and trihydroxylated bile acids from their ketones and unconjugated versions to give 96 bile acids (Figure 4a). A univariate analysis at Step 1 of bile acid annotations showed a higher abundance of almost all bile acids in the carnivores compared to herbivores and omnivores combined (Figure 4b). As we move down the MassQL filtering tree to Step 4, we can now hypothesize the regio- and, in many cases, the stereochemistry of their hydroxyl positions. With the current implementation of the tree, we can predict the bile acid cores of 49 MS/MS spectra (Figure 4a). The highest number of spectral matches that could be assigned are dihydroxylated bile acids, symbolized here as (OH)2 (Figure 4c). These were categorized into individual bins of 3,12α-(OH)2, 3,7-(OH)2; 3,6-(OH)2; 3,12β-(OH)2; 7,12β-(OH)2 and 3,6-(OH)2 isomers following the application of the dihydroxy isomer filtering tree.
Conclusion
In conclusion, this study introduces the concept of using a MassQL MS/MS fragmentation-based filtering tree approach to enable precise regio- and stereochemical differentiation of bile acid isomers, which were subsequently validated against reference standards. The method expands the ability to annotate bile acids and has led to the discovery of bile acid deoxycholyl-N-acetyl-putrescine, which had previously been detected in the feces of carnivores but not structurally characterized. By making annotated MS/MS spectra publicly available and enabling the MassQL queries, this approach enhances bile acid analysis, offering innovative tools for exploring their metabolic roles and biological significance, including retro-analysis of regio- and stereochemical isomer variations in existing public LC-MS/MS data.