Cerebrospinal Fluid Proteome Reveals Dysregulation of Lysosomal and Axonal Proteins in Neurosyphilis
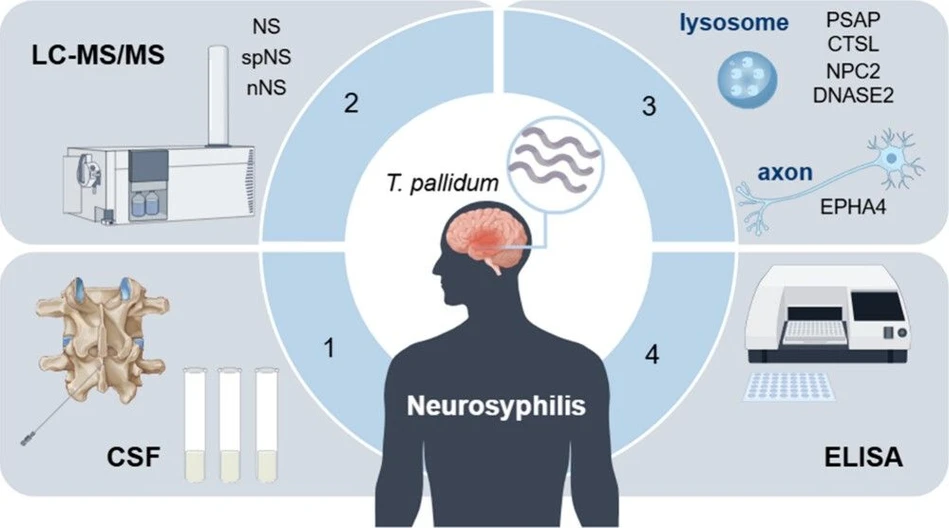
J. Proteome Res. 2026, 25, 2, 713–722: Graphical abstract
This study investigates the pathogenesis of neurosyphilis through cerebrospinal fluid proteomics using LC–MS/MS and ELISA validation. Comparative analysis identified 234 differentially expressed proteins, with major alterations linked to lysosomal dysfunction, axon guidance, and neurodegeneration pathways.
Key lysosomal proteins, including PSAP and DNASE2, together with the axonal protein EPHA4, were significantly downregulated in neurosyphilis patients. These proteins showed potential as biomarkers for neurodegeneration and may provide new insights into disease mechanisms, diagnostics, and therapeutic targets.
The original article
Cerebrospinal Fluid Proteome Reveals Dysregulation of Lysosomal and Axonal Proteins in Neurosyphilis
Huayang Tang*, Junjie Hu, Yifan Wu, Jinping Gao, Wenjun Wang, Xiaodong Zheng, Ran Zhang, Bo Liang, Fusheng Zhou, Ze Guo*
J. Proteome Res. 2026, 25, 2, 713–722
https://doi.org/10.1021/acs.jproteome.5c00689
licensed under CC-BY 4.0
Selected sections from the article follow. Formats and hyperlinks were adapted from the original.
Neurosyphilis (NS) is a group of clinical syndromes characterized by neuroinflammation caused by Treponema pallidum (T. pallidum), which occurs in all stages of syphilis. (1,2) NS is classified based on clinical presentation as either asymptomatic (ANS) or symptomatic (SNS), and based on disease stage as either early (occurring 1–2 years after primary infection) or late. (1) T. pallidum can cause local nonspecific and central nervous system (CNS) inflammation, leading to possible degenerative changes. (3) The immune response of CNS in neurosyphilis is comparable to that observed in other neuroinflammatory diseases. Consequently, the presence of neurological symptoms may not reliably indicate active CNS inflammation in syphilis patients. (3) A cerebrospinal fluid (CSF) proteomic study identified three biomarkers (SEMA7A, SERPINA3, and ITIH4) that contribute to NS and have potential implications for diagnosis and therapeutic strategies. (4) Through integrated proteomics and single-cell transcriptomics analyses, Toll-like/NF-κB signaling was identified as the key pathway involved in CNS damage caused by T. pallidum. (5) Despite an increasing understanding of the biomarkers and possible neuroinflammatory damage mechanisms of neurosyphilis, the mechanisms of neurodegeneration caused by T. pallidum infection remain elusive. (6)
Patients with neurosyphilis exhibit stage-specific profiles of soluble factors in their CSF, which are linked to both immune status and the extent of neuronal damage. (7) In this study, we analyzed differentially expressed proteins (DEPs) in the CSF proteomes from 28 individuals, including neurosyphilis (NS), suspected neurosyphilis (spNS) and non-neurosyphilis (nNS) individuals, by LC–MS/MS and analyzed a public data set of the neurosyphilis CSF proteome. Validation was then performed by ELISA in an independent group comprising 110 samples. The results revealed that neuroinflammation by T. pallidum infection results in dysregulation of the key proteins in lysosomal and axonal pathway, which suggesting a potential contribution to neurodegeneration in neurosyphilis.
Materials and Methods
LC–MS/MS
The tryptic peptides were dissolved in 0.1% formic acid (solvent A) and loaded onto a custom-made reversed-phase column (15 cm × 75 μm). Using an EASY-nLC 1000 UPLC system, a gradient was run at 400 nL/min as follows: 6% to 23% solvent B (0.1% formic acid in 98% acetonitrile) over 26 min, 23% to 35% in 8 min, up to 80% in 3 min, and held at 80% for 3 min. Peptides were ionized via an NSI source at 2.0 kV and analyzed online by a Q Exactive Plus mass spectrometer. Full scans (m/z 350–1800) were acquired in the Orbitrap at 70,000 resolution. The top precursors were selected for MS/MS with an NCE of 28, and fragments were detected at 17,500 resolution. A data-dependent method cycled between one MS scan and 20 MS/MS scans with a 15.0 s dynamic exclusion. AGC was set at 5E4. MS/MS data were processed with MaxQuant (v.1.5.2.8) using Trypsin/P as the enzyme (up to 2 missed cleavages). Precursor mass tolerance was 20 ppm (first search) and 5 ppm (main search); fragment mass tolerance was 0.02 Da. Fixed modification: carbamidomethyl (C); variable modification: oxidation (M). The FDR threshold was <1%, and the minimum peptide score was >40.
ELISA
CSF levels of host proteins were quantified using commercial ELISA kits according to the manufacturers’ protocols. Proteins analyzed included PSAP (OKEH07124, Aviva, USA), CTSL (abx150963, Abbexa, UK), NPC2 (A-QEK07857, Biogradetech, USA), and DNASE2 (abx386953, Abbexa, UK). Briefly, 100 μL of CSF was added to precoated plates and incubated at 37 °C for 2 h. After washing, 100 μL of detection antibody was added to each well and incubated at 37 °C for another 2 h. Following a wash, 100 μL of streptavidin-HRP working solution was added and incubated at room temperature for 20 min in the dark. The reaction was stopped with 50 μL of stop solution, and the optical density was measured at 450 nm.
Results
Downregulation of Key Lysosomal Proteins in CSF from Patients With Neurosyphilis
By analyzing the DEPs in NS vs nNS, we observed a significant enrichment of downregulated proteins in the lysosome pathway (P = 0.007) (Figure 2A), with 9 lysosomal proteins showing significant downregulation (Padj < 0.05) in the NS group (Table S4). To validate this finding, we examined the public CSF proteomics data set of neurosyphilis (PXD033034) and identified 18 proteins in the lysosomal pathway that were significantly downregulated in patients with NS (Padj < 0.05) (Table S4). PPI network analysis determined the top 10 hub genes within the lysosome pathway (Figure 2B). A Venn diagram revealed 3 proteins (PSAP, CTSL, and NPC2) shared across our current data, the public data set (PXD033034) and the top 10 hub genes (Figure 2C). The mass spectrometry results indicated that the levels of PSAP (Padj = 0.043), CTSL (Padj = 0.028) and NPC2 (Padj = 0.013) were significantly lower in the NS group compared to the nNS group (Figure 2D). The PXD033034 data set revealed consistent changes in the PSAP, CTSL, and NPC2 levels in the NS group (Padj < 0.05) (Figure S1 and Table S4). In addition, associations between the lysosomal protein DNASE2 and NS were observed in both the present study (Padj = 0.003) (Figure 2D) and the PXD033034 data set (Padj < 0.05) (Figure S1 and Table S4), with subgroup analysis revealing a significantly lower expression level of DNASE2 in the SNS group compared to the ANS group (P = 0.035) (Figure 2D and Table S3). Further validation by ELISA (Table S5) in an independent group (Table 1) confirmed significantly downregulated CSF levels of PSAP (P < 0.0001), CTSL (P < 0.0001), NPC2 (P = 0.0024) and DNASE2 (P < 0.0001) in patients with NS (n = 21) than in nNS individuals (n = 56) (Figure 2E) (Table S6), as well as decreased in the SNS subgroup (P < 0.05) (Figure S2 and Table S6). Receiver operating characteristic (ROC) curve analysis supported PSAP (AUC = 0.91, 95% CI: 0.83–0.99) and DNASE2 (AUC = 0.95, 95% CI: 0.91–0.99) for distinguishing NS from nNS and as potential biomarkers for assessing neurodegeneration in NS (Figure 2F). These results indicate that T. pallidum infection may lead to lysosomal dysfunction in CNS.
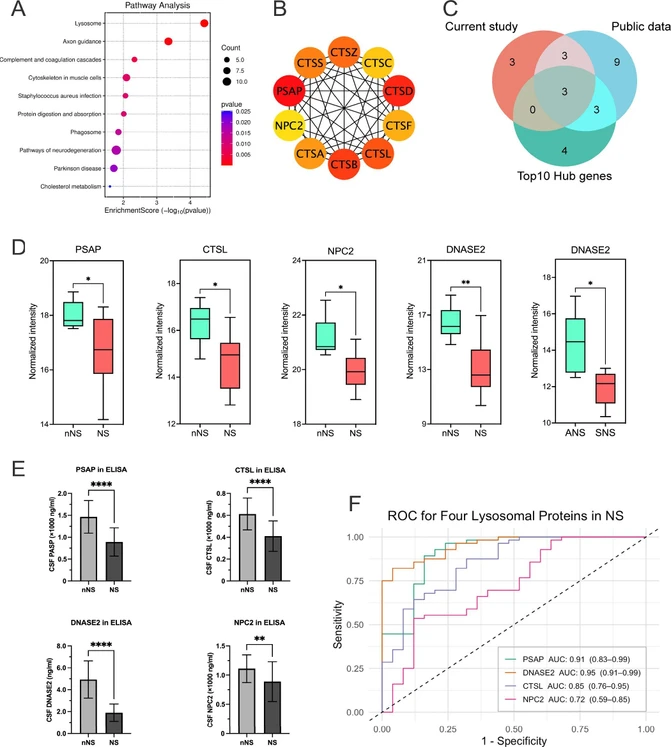 J. Proteome Res. 2026, 25, 2, 713–722: Figure 2. Downregulation of key lysosomal proteins in the cerebrospinal fluid of neurosyphilis patients. (a) KEGG pathway analysis demonstrated that downregulated proteins in patients with NS were significantly enriched in the lysosome pathway, as well as in axon guidance and neurodegeneration. (b) The top 10 hub genes involved in the lysosome pathway were identified using the STRING network data set and PPI analysis. (c) A Venn diagram revealed 3 proteins (PSAP, CTSL, and NPC2) shared across our current data, the public data set (PXD033034) and the top 10 hub genes. (d) The CSF levels of PSAP (Padj = 0.043), CTSL (Padj = 0.028), DNASE2 (Padj = 0.003) and NPC2 (Padj = 0.013) were significantly decreased in the NS group than in the nNS group. Subgroup analysis found a significantly lower expression level of DNASE2 in the SNS group than in the ANS group (P = 0.035). (e) PSAP, DNASE2, CTSL and NPC2 levels in CSF were tested by ELISA in the nNS (n = 56) and NS (n = 21) groups. Compared with those in the nNS group, significantly lower PSAP (P < 0.0001), CTSL (P < 0.0001), DNASE2 (P < 0.0001) and NPC2 (P = 0.0024) levels were detected in the NS group. (f) ROC curve analysis for NS revealed that PSAP had an AUC = 0.91 (95% CI: 0.83–0.99), DNASE2 had an AUC = 0.95 (95% CI: 0.91–0.99), CTSL had an AUC = 0.85 (95% CI: 0.76–0.95), and NPC2 had an AUC = 0.72 (95% CI: 0.59–0.85). NS: neurosyphilis; nNS: non-neurosyphilis; ANS: asymptomatic neurosyphilis; SNS: symptomatic neurosyphilis. ROC: receiver operating characteristic; AUC: area under the curve. * P < 0.05, ** P < 0.01, ****P < 0.0001.
J. Proteome Res. 2026, 25, 2, 713–722: Figure 2. Downregulation of key lysosomal proteins in the cerebrospinal fluid of neurosyphilis patients. (a) KEGG pathway analysis demonstrated that downregulated proteins in patients with NS were significantly enriched in the lysosome pathway, as well as in axon guidance and neurodegeneration. (b) The top 10 hub genes involved in the lysosome pathway were identified using the STRING network data set and PPI analysis. (c) A Venn diagram revealed 3 proteins (PSAP, CTSL, and NPC2) shared across our current data, the public data set (PXD033034) and the top 10 hub genes. (d) The CSF levels of PSAP (Padj = 0.043), CTSL (Padj = 0.028), DNASE2 (Padj = 0.003) and NPC2 (Padj = 0.013) were significantly decreased in the NS group than in the nNS group. Subgroup analysis found a significantly lower expression level of DNASE2 in the SNS group than in the ANS group (P = 0.035). (e) PSAP, DNASE2, CTSL and NPC2 levels in CSF were tested by ELISA in the nNS (n = 56) and NS (n = 21) groups. Compared with those in the nNS group, significantly lower PSAP (P < 0.0001), CTSL (P < 0.0001), DNASE2 (P < 0.0001) and NPC2 (P = 0.0024) levels were detected in the NS group. (f) ROC curve analysis for NS revealed that PSAP had an AUC = 0.91 (95% CI: 0.83–0.99), DNASE2 had an AUC = 0.95 (95% CI: 0.91–0.99), CTSL had an AUC = 0.85 (95% CI: 0.76–0.95), and NPC2 had an AUC = 0.72 (95% CI: 0.59–0.85). NS: neurosyphilis; nNS: non-neurosyphilis; ANS: asymptomatic neurosyphilis; SNS: symptomatic neurosyphilis. ROC: receiver operating characteristic; AUC: area under the curve. * P < 0.05, ** P < 0.01, ****P < 0.0001.
Downregulation of Key Axonal Proteins Correlated with Lysosomal Protein Reduction in Neurosyphilis CSF
Lysosomes play crucial roles in neuronal health by maintaining axonostasis. (10) The enrichment analysis indicated that the downregulated proteins in NS were implicated in the biological processes of axonostasis (such as axon development and axonogenesis) (Figure 1F) and the pathway of axon guidance (Figure 2A). Our proteomic data revealed a total of 24 downregulated (Padj < 0.05) axonal proteins and 1 upregulated (Padj < 0.05) axonal protein in the CSF of patients with NS (Table S9). Furthermore, the PXD033034 data set revealed 55 downregulated (Padj < 0.05) and 4 upregulated (Padj < 0.05) axonal proteins in NS (Table S9). The top 10 hub genes involved in the biological processes of axon development and axonogenesis were identified by PPI network analysis (Figure 4A). Considering the overlap of our current data, the PXD033034 data set and the top 10 hub genes (Figure 4B), one key protein, EPHA4, was identified across 3 data sets, with significant downregulation in our current data (Padj = 0.011) and in the PXD033034 data set (Padj = 0.028) (Figure 4C). Correlation analysis further revealed a positive correlation between the levels of PSAP and EPHA4 (r = i 0.83, P < 0.05) (Figure 4D). Reduced levels of PSAP and EPHA4 in CSF have been implicated in neurodegenerative diseases, including Alzheimer’s disease (AD) and Parkinson’s disease (PD). (11,12) Our findings suggest that coordinated dysregulation of lysosomal and axonal proteins may contribute to neurodegeneration in NS.
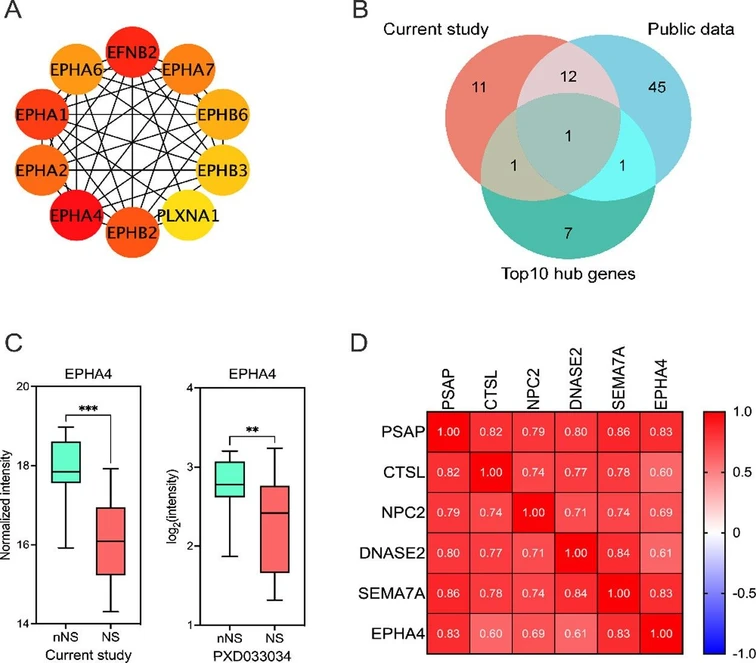 J. Proteome Res. 2026, 25, 2, 713–722: Figure 4. Key axonal protein downregulated in CSF from patients with neurosyphilis. (a) The top 10 hub genes involved in axonostasis were identified by the STRING network data set and PPI analysis. (b) A Venn diagram revealed 1 protein (EPHA4) shared across DEPs related to axonostasis in our current data and in the public data set (PXD033034) and the top 10 hub genes. (c) EPHA4 was significantly downregulated both in our present study (Padj = 0.011) and in the PXD033034 data set (Padj = 0.028). (d) Correlation analysis demonstrated that the EPHA4 level was significantly positively correlated (P < 0.05) with the levels of PSAP (r = 0.83), CTSL (r = 0.6), NPC2 (r = 0.69), DNASE2 (r = 0.61) and SEMA7A (r = 0.83). NS: neurosyphilis; nNS: non-neurosyphilis. * P < 0.05.
J. Proteome Res. 2026, 25, 2, 713–722: Figure 4. Key axonal protein downregulated in CSF from patients with neurosyphilis. (a) The top 10 hub genes involved in axonostasis were identified by the STRING network data set and PPI analysis. (b) A Venn diagram revealed 1 protein (EPHA4) shared across DEPs related to axonostasis in our current data and in the public data set (PXD033034) and the top 10 hub genes. (c) EPHA4 was significantly downregulated both in our present study (Padj = 0.011) and in the PXD033034 data set (Padj = 0.028). (d) Correlation analysis demonstrated that the EPHA4 level was significantly positively correlated (P < 0.05) with the levels of PSAP (r = 0.83), CTSL (r = 0.6), NPC2 (r = 0.69), DNASE2 (r = 0.61) and SEMA7A (r = 0.83). NS: neurosyphilis; nNS: non-neurosyphilis. * P < 0.05.
Conclusions
The present study reveals the associations of dysregulated proteins in lysosomal and axonal pathways with neurosyphilis, and potential evidence of shared pathophysiological mechanisms between neuroinflammation caused by T. pallidum infection and neurodegenerative diseases.
However, this study has certain limitations. First, the use of ELISA to validate a limited number of proteins has certain limitations, such as the inherent cross-reactivity of ELISA. In the future, we can optimize the validation experiments by adding techniques such as Western blotting for verification. Second, this study uses a cross-sectional sampling method for research, which limits our ability to infer disease progression and treatment responses. In the future, we will conduct follow-up on the subjects, continue collecting cerebrospinal fluid samples after treatment, and observe whether the protein dysregulation is reversible after treatment. For spNS patients, follow-up should also be conducted to determine if they have transformed into NS, which will be very valuable for validating the predictive utility of our biomarkers. Finally, although multiple discovery and validation cohort studies have been conducted, we currently lack direct mechanistic evidence. Future work will investigate whether exposing neural cell coculture models to outer-membrane antigens of T. pallidum reproduces the lysosome and axon alterations observed in CSF, clarifying their role in neural damage of neurosyphilis and potential biomarkers for diagnosis and treatment of neurodegeneration in neurosyphilis.