LCMS-Net: Deep Learning for Raw High Resolution Mass Spectrometry Data Applied to Forensic Cause-of-Death Screening
Anal. Chem. 2026, 98, 9, 6589–6597: Figure 1. Overview of LCMS-Net applied to CoD screening. Binning is used to reduce the size of the raw LC-HRMS data and to ensure equally shaped input matrices. Afterward, the input matrices are processed by an ensemble of 1D-CNNs consisting of a depthwise 1D convolutional layer, followed by max pooling and spatial dropout. The results of the CNNs are flattened and dense layers are used to predict the class labels of the input data.
This study introduces LCMS-Net, a deep learning model designed for direct analysis of raw LC-HRMS data without the need for manual preprocessing. By modeling spatial features of the data, the approach improves reproducibility and detection performance in untargeted metabolomics workflows.
LCMS-Net demonstrated improved performance in forensic cause-of-death screening and colon cancer detection, outperforming existing methods while reducing batch effects across instruments. Its simplified architecture and lack of pretraining requirements enable faster and more efficient data analysis, highlighting its potential for routine metabolomics and forensic applications.
The original article
LCMS-Net: Deep Learning for Raw High Resolution Mass Spectrometry Data Applied to Forensic Cause-of-Death Screening
Lisa M. Menacher, Liam J. Ward, Fredrik Heintz, Henrik Green, and Oleg Sysoev*
Anal. Chem. 2026, 98, 9, 6589–6597
https://doi.org/10.1021/acs.analchem.5c05404
licensed under CC-BY 4.0
Selected sections from the article follow. Formats and hyperlinks were adapted from the original.
Metabolomics is the study of all low-molecular-weight substances in a biological specimen. (1) As this is closely linked to cellular functions and biochemical processes, it offers a comprehensive view of the current physiological state of an organism. This makes metabolomics a promising tool for a broad range of applications, including diagnostics, precision medicine, and toxicology. (1−3) However, large-scale large-scale liquid chromatography-high resolution mass spectrometry (LC-HRMS) studies are difficult due to the complex and labor-intensive data preprocessing. This typically involves peak picking, peak alignment, gap filling, and normalization, to simplify the data structure, extract biologically relevant features, and improve the interpretability for subsequent analyses. (4−7) Although tools like XCMS or MZmine offer widely used implementations of this workflow, the preprocessing of LC-HRMS data remains challenging, for a number of reasons: (1) The preprocessing of LC-HRMS data can be time-consuming due to computational constraints for large datasets. (4,8,9) (2) Domain knowledge is necessary to select suitable parameters for the used preprocessing software, and thus to obtain reliable features. (4,7,8) (3) The reproducibility of the extracted features is often limited across software libraries, especially for low-abundant metabolites. (4,8) (4) Preprocessing tools may fail to detect some metabolites entirely, resulting in the loss of potentially important information. (4)
In the past few years, deep learning models have emerged as promising alternatives to the traditional preprocessing workflow for LC-HRMS data. While many publications in this field have been focused on enhancing a particular step of the preprocessing, such as peak identification or peak alignment, end-to-end deep learning has also gained popularity. (9,10) In end-to-end deep learning, a model is trained to extract useful information directly from raw data without explicit feature engineering. Often, this is less time-consuming than conventional methods and improves the predictive accuracy of the downstream task. This was demonstrated by Cadow et al., who converted raw MS data from prostate biopsies into pseudoimages through binning. They then applied pretrained deep learning models for image processing to extract feature vectors from the pseudoimages and trained a classifier to discriminate between tumor and healthy cases. (11) A similar approach was also used by Shen et al. to predict the gestational age of pregnant women. To address data scarcity, they proposed a data augmentation strategy that simulates retention time (RT) drifts during the data acquisition. As a result, several pseudoimages can be produced from a single LC-HRMS sample, which substantially increases the size of the training set. (12) Furthermore, Wang et al. developed an end-to-end deep learning model to detect esophageal squamous cell carcinoma based on raw LC-HRMS data. By decomposing a sample into multiple tiles instead of just one down-scaled pseudoimage, their approach preserves more details of the raw LC-HRMS data. (13) Most recently, Deng et al. used an ensemble of pretrained image classification models for lung cancer detection. Their approach enables more robust predictions for multiclass classification tasks. Furthermore, model explainability was addressed through the analysis of contributing factors. (14) Although, these studies collectively highlight the potential of end-to-end deep learning for metabolomics-based analyses, they also show its limitations. For example, the use of pretrained architectures for image classification may be flawed due to the substantially different topology of LC-HRMS data and natural images. Natural images typically contain hierarchically structured objects with geometrical properties such as size or patterns while objects in LC-HRMS pseudoimages are essentially represented as one-dimensional peaks along the retention time axis and are sharply localized in m/z-value. (6,15) Existing deep learning methods for metabolomics primarily use 2D-convolutions and thus do not take into account such spatial properties of LC-HRMS data. Consequently, there remains a need for more robust methods that are specifically tailored to LC-HRMS data. This is particularly relevant for real-world applications, where batch effects and diverse study populations are common and difficult to mitigate.
This study presents LCMS-Net, a fully automated end-to-end deep learning model for the analysis of LC-HRMS data. By directly processing the raw data and taking into account its spatial properties by using 1D-convolutions, LCMS-Net mitigates the need for time-intensive, manual feature engineering and improves the classification performance of downstream tasks. This is demonstrated through comprehensive benchmarking for two different applications, forensic CoD screening and colon cancer detection. To the best of our knowledge, it is the first time end-to-end deep learning is applied to post-mortem metabolomics and in the context of forensic science. Furthermore, LCMS-Net achieves robust results even in the presence of severe batch effects, as shown by training and evaluating the CoD screening model with data from different measurement instruments. Lastly, LCMS-Net does not rely on large pretrained deep learning models, and thus requires less computational resources than previous end-to-end learning approaches for the analysis of LC-HRMS data.
Methods
Cause-of-Death Screening Dataset
The cause-of-death screening study was approved by the Swedish Ethical Review Authority (Dnr 2019–04530 & Dnr 2025–0249–02). All samples were collected from autopsy cases admitted to the Swedish National Board of Forensic Medicine between July 2017 and December 2024, which had undergone routine toxicological screening via HRMS. Screening was performed from femoral whole blood samples, in accordance with standardized procedures reported previously. (32) Sample separation was performed using gradient elution on a C18 column (150 mm × 2.1 mm, 1.8 μm; Waters Acquity HSS T3 column, Water Sverige AB, Sweden). MS-data was collected in positive mode and the total acquisition time for each sample was 12 min. Samples admitted between July 2017 and November 2020 were run using an Agilent 6540 Q-TOF system in MS-mode only, and samples admitted afterward using an Agilent 6546 Q-TOF system with data-dependent acquisition mode (autoMSMS). Aside from these differences in acquisition strategy, source settings and TOF parameters (e.g., gas temperatures, voltages, and mass ranges) were kept consistent between platforms. Henceforth, we will refer to the former as Dataset A and the latter as Dataset B for clarity.
Dataset A was primarily used for the implementation and evaluation of LCMS-Net and includes 4282 autopsy cases processed in over 641 analytical runs. All cases belong to one of the following five CoD groups: acidosis (n = 100), drug intoxication (n = 1385), ischemic heart disease (IHD) (n = 1362), hanging (n = 1200), and pneumonia (n = 235). A detail summary of the inclusion criteria and study population was previously described by Ward et al. (26) Dataset B was used to evaluate the sensitivity of LCMS-Net to batch effects and includes additional 5485 autopsy cases with the same inclusion criteria and CoD groups as Dataset A.
Overview of LCMS-Net
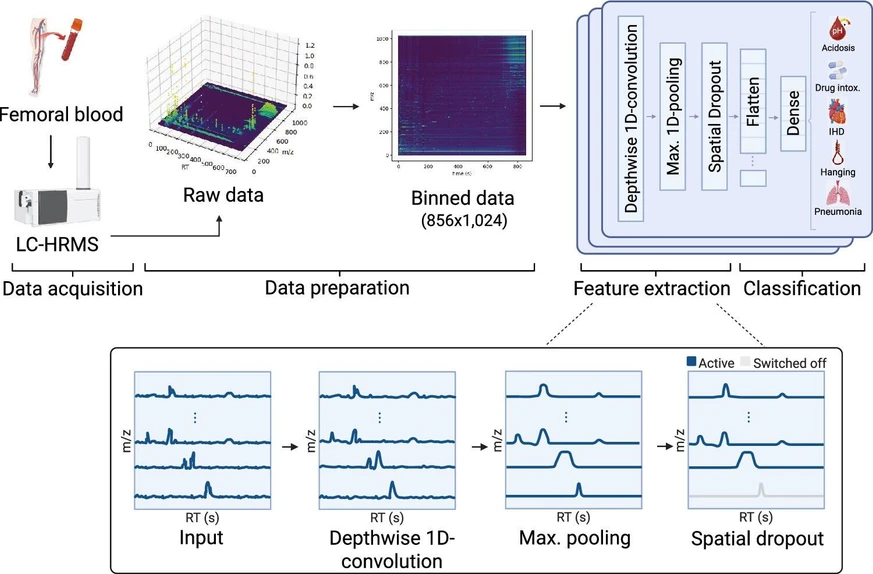
LCMS-Net is a end-to-end deep learning model for the analysis of raw LC-HRMS data. First, a data preparation pipeline transforms the raw LC-HRMS data into pseudoimages. Afterward, convolutional neural networks (CNNs) specifically designed to take the spatial properties of LC-HRMS data into account are used to extract relevant features for the downstream task. Figure 1 shows an overview of this workflow for CoD screening, as well as the architecture of LCMS-Net.
 Anal. Chem. 2026, 98, 9, 6589–6597: Figure 1. Overview of LCMS-Net applied to CoD screening. Binning is used to reduce the size of the raw LC-HRMS data and to ensure equally shaped input matrices. Afterward, the input matrices are processed by an ensemble of 1D-CNNs consisting of a depthwise 1D convolutional layer, followed by max pooling and spatial dropout. The results of the CNNs are flattened and dense layers are used to predict the class labels of the input data.
Anal. Chem. 2026, 98, 9, 6589–6597: Figure 1. Overview of LCMS-Net applied to CoD screening. Binning is used to reduce the size of the raw LC-HRMS data and to ensure equally shaped input matrices. Afterward, the input matrices are processed by an ensemble of 1D-CNNs consisting of a depthwise 1D convolutional layer, followed by max pooling and spatial dropout. The results of the CNNs are flattened and dense layers are used to predict the class labels of the input data.
Results and Discussion
Evaluation of Prediction Performance for CoD Screening
First, LCMS-Net was evaluated on the CoD screening dataset (Dataset A). Figure 2 shows the results of the model comparison for the CoD screening dataset. With an average accuracy of 70.8%, LCMS-Net outperformed all four benchmark models trained on preprocessed LC-HRMS data. In comparison, our previously published OPLS-DA model for CoD screening achieved only 65.0% accuracy (p-value = 0.0010). Furthermore, LCMS-Net also achieved a better F1-score and sensitivity than these models, while maintaining a high specificity score. The deep learning benchmark model, DeepMSProfiler, achieved a similar accuracy to LCMS-Net. However, LCMS-Net outperformed DeepMSProfiler in terms of both F1-score (p-value = 0.0122) and sensitivity (p-value = 0.0010). This indicates that our method maintains a better balance between true positives and false positives across the five CoD groups, which is especially important for high-stakes applications like forensic science. It should be noted that traditional cross-validation was used for the estimation of p-values, as our purpose was to compare models with fixed hyperparameters. When instead comparing frameworks (e.g., RF vs SVM), nested cross-validation would be more appropriate.
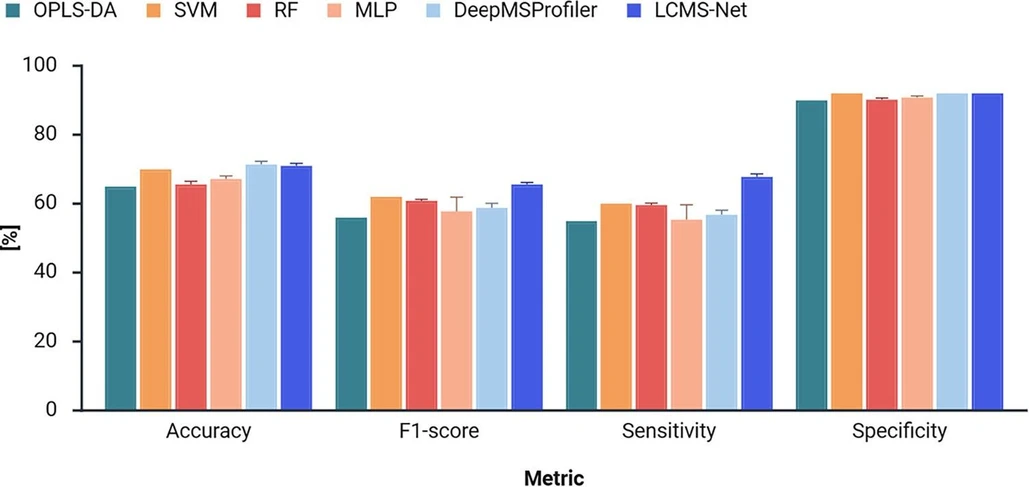 Anal. Chem. 2026, 98, 9, 6589–6597: Figure 2. Evaluation of CoD screening prediction performance of LCMS-Net in comparison to benchmark models. The prediction performance of a classifier is measured by accuracy, macro F1-score, macro sensitivity, and macro specificity over five model runs. The error bars represent the standard deviations between model runs.
Anal. Chem. 2026, 98, 9, 6589–6597: Figure 2. Evaluation of CoD screening prediction performance of LCMS-Net in comparison to benchmark models. The prediction performance of a classifier is measured by accuracy, macro F1-score, macro sensitivity, and macro specificity over five model runs. The error bars represent the standard deviations between model runs.
Notably, LCMS-Net achieves its high classification performance without pretraining on natural images. This shows that although transfer learning is often useful, it may fail if the source (e.g., natural images) and target data distribution (e.g., LC-HRMS data) are too different. Such domain mismatch can lead to negative transfer, where features from a pretrained model negatively impact the classification performance of the downstream task, resulting in a lower accuracy than training a simpler model from scratch. (42) By employing an architecture tailored to LC-HRMS data, LCMS-Net is also computationally more efficient, with fewer than 500,000 trainable parameters compared to over 7 million trainable parameters in DeepMSProfiler. (14) This makes our method accessible to researchers with limited computational resources.
Application for Colon Cancer Detection
Lastly, we have tested the potential of LCMS-Net in another application domain, by assessing its prediction performance on the CRC dataset. The cancer detection task has been previously used by Deng et al. to show that DeepMSProfiler can be successfully applied to different application areas, making it a suitable benchmark task. We trained a separate instance of LCMS-Net for the model evaluation on the CRC dataset due to differences in sample preparation, analytical protocols, and instrument setup compared to the CoD screening dataset. The hyperparameter settings remained unchanged, as we assumed similar structural properties for both datasets.
LCMS-Net outperforms DeepMSProfiler with an F1-score of 97.3% compared to 95.5% on the colon cancer detection task. Furthermore, sensitivity and specificity improvements of 2.8% were observed. These results underline the relevance of LCMS-Net for other application areas than CoD screening. A detailed overview of the evaluation metrics for the colon cancer prediction can be found in Supporting Table 7.
Conclusion
Current preprocessing workflows for untargeted metabolomics are often time-consuming, require extensive domain knowledge, lack reproducibility, or fail to detect some metabolites entirely. LCMS-Net offers an alternative end-to-end approach by operating directly on the raw LC-HRMS data and explicitly modeling its spatial properties. The deep learning model enables faster processing and does not require manual parameter selection. Furthermore, it has been shown through two case studies, CoD screening and cancer detection, that our method extracts a higher amount of relevant information from the raw data and thus outperforms existing models for the analysis of metabolomics data. Specifically, LCMS-Net achieves an average F1-score of 65.5% for CoD screening and 97.3% for colon cancer detection. The previous state-of-the-art models achieved 56.1% (OPLS-DA) and 95.5% (DeepMSProfiler) respectively. The prediction performance of LCMS-Net is consistent even when applied to data from different measurement instruments than those used for the training of the deep learning model, showing its robustness toward batch effects. However, the interpretability of LCMS-Net remains limited and future development is needed to assess which metabolites are learned by the model and which remain undetected. Despite this, LCMS-Net opens up new opportunities for large-scale analyses of LC-HRMS data, potentially driving innovation across numerous application domains.