Investigating the Effect of Isoelectric Points on the Gas-Phase Stability of Native-like Proteins Analyzed in Positive- versus Negative-Ion Mode by IMS-MS
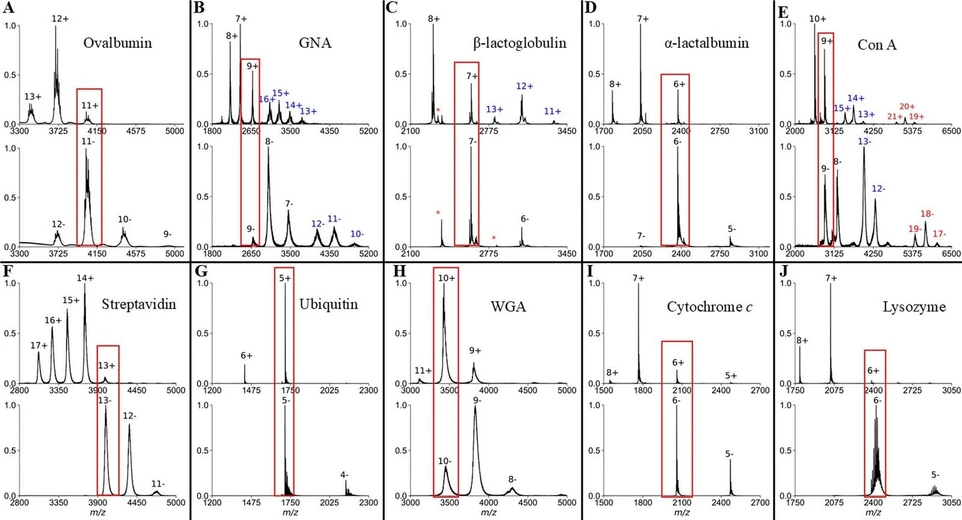
Anal. Chem. 2026, 98, 6, 4518–4527: Figure 1. Representative mass spectra in +ESI (top) and −ESI (bottom) for (A) ovalbumin, (B) GNA, (C) β-lactoglobulin, (D) α-lactalbumin, (E) con A, (F) streptavidin, (G) ubiquitin, (H) WGA, (I) cytochrome c, and (J) lysozyme. Proteins that have blue or red charge states indicate additional oligomeric states that were detected. Outlines in red boxes are overlapping charge states that are compared herein. *Indicates contaminant peaks.
This study investigates how ionization polarity influences the gas-phase stability of native-like proteins analyzed by ion mobility spectrometry–mass spectrometry. While proteins generally retain folded structures in both positive and negative electrospray ionization modes, significant differences in unfolding behavior were observed.
Proteins exhibited greater gas-phase stability when analyzed in the ionization polarity corresponding to their solution-phase charge, requiring higher energy for unfolding. These findings emphasize the importance of considering protein isoelectric points and ionization polarity in native IMS-MS experiments to ensure accurate structural characterization.
The original article
Investigating the Effect of Isoelectric Points on the Gas-Phase Stability of Native-like Proteins Analyzed in Positive- versus Negative-Ion Mode by IMS-MS
Alexis N. Edwards, Madeline G. Bannon, Michael S. Cordes, and Elyssia S. Gallagher*
Anal. Chem. 2026, 98, 6, 4518–4527
https://doi.org/10.1021/acs.analchem.5c04295
licensed under CC-BY 4.0
Selected sections from the article follow. Formats and hyperlinks were adapted from the original.
In native mass spectrometry (nMS), the native-like structures of proteins are maintained in the gas phase by preserving the secondary, tertiary, and quaternary structures. (1−7) The preservation of native-like protein structures is primarily accomplished via the use of electrospray ionization (ESI), a soft ionization method that forms a distribution of multiply charged protein ions. These multiply charged ions can then be investigated to characterize stoichiometry, oligomeric composition, and ligand binding. (8−10)
During ESI, a potential is applied to the sample-containing solution. In an aqueous spray solvent, H3O+ and OH– are produced in positive-ion mode (+ESI) and negative-ion mode (−ESI), respectively. (11) The buildup of like-charges within the solution results in the formation of a Taylor Cone at the capillary tip, which ejects charged droplets that travel toward the inlet of the mass spectrometer. Droplets undergo evaporation and fission until complete analyte desolvation, leading to the release of folded, gas-phase protein ions. As the droplets shrink, H3O+ or OH– can become concentrated, causing the pH of the droplet to decrease or increase, respectively. Shifts in pH can denature proteins. Therefore, to maintain native-like protein structures, nMS is conducted with solutions containing volatile salts, usually ammonium acetate, that provide ionic strength and minimize pH changes compared to spraying from pure water. (11,12) Droplet pH in ESI is influenced by flow rate, solvent composition, and applied voltage, (13) but often approaches values near the pKa of acetate (4.76) or ammonium (9.25) in +ESI or −ESI, (12,14) respectively. During ESI, changes in overall protein charge take place via proton transfer at chargeable amino acid residues, resulting in the formation of protonated or deprotonated protein ions with the charged sites being dependent on the relative proton affinities of ammonium, acetate, and chargeable amino acids as proteins near complete desolvation. (15−19)
Ion mobility spectrometry (IMS) measures an ion’s mobility through an inert gas under the influence of an electric field. Thus, mobility is directly influenced by the ion’s size, shape, and charge. (20) A protein’s mobility can be used to estimate its structure by calculating an average momentum-transfer collisional cross section (CCS). In addition to structural characterization, IMS-MS can investigate the gas-phase stability of protein complexes when paired with collision-induced unfolding (CIU). Specifically, in CIU experiments, protein ions are accelerated into a cell filled with inert gas molecules and undergo multiple collisions with higher frequency and energy relative to IMS. This converts kinetic energy to internal, vibrational energy, disrupting noncovalent interactions and inducing unfolding of the protein ions. (21) Unfolding alters the protein structure, which can then be quantified by monitoring the protein’s mobility during subsequent IMS-MS measurement. (22) Prior work with mutant proteins has shown that proteins with similar intramolecular interactions have similar CIU fingerprints, while proteins with different intramolecular interactions have different CIU fingerprints. (23) Therefore, gas-phase protein stability is correlated with intramolecular interactions.
Past native IMS-MS work has primarily been conducted in +ESI given that -ESI has the reputation of being unreliable for native-like protein analysis due to the occurrence of corona discharge, (24) and charge-state distributions that may not be representative of folded proteins. (25) In recent years, −ESI has been increasingly used in CIU and IMS-MS, with recent work demonstrating that −ESI-MS is capable of discerning denatured and native-like protein structures. (26,27) Prior work showed that charge manipulation of folded proteins, such as charge reduction, to achieve similar absolute charge states in both ion modes, results in CCS values of protein cations that are comparable to their anion counterparts. (26,28) This suggests that gas-phase structures are weakly dependent on protein charge and ionization polarity. (23,26,28) Yet, Hong et al. showed that, in the absence of charge reduction, the gas-phase stability of two proteins (avidin and β-lactoglobulin) is sensitive to ionization polarity, highlighting that gas-phase stability is not consistent between +ESI and −ESI. (29)
Materials and Methods
Native IMS-MS Analysis
Glass capillaries were pulled using a P-1000 micropipette puller (Sutter Instrument Company, Novato, CA). Samples and calibrants were sprayed from pulled capillaries (inner diameter ∼1 μm) and analyzed with a Synapt G2-S High-Definition MS (Waters Corporation, Milford, MA). The trap and transfer cells contained Ar, the helium cell contained He, and the IMS cell contained N2. IMS wave velocity was 350 m/s, while IMS wave height was either 7 or 9 V, depending on the protein size (see Table S2). Proteins with larger mass required a wave height of 9 V to ensure that all the ions migrated out of the IMS cell prior to the next ion injection. Mass spectra were collected and signal averaged over 30 s. Additional instrument details can be found in the Supporting Information (Table S2).
To limit systematic error due to repositioning of the ESI tip between runs, (48) the XYZ-stage position was optimized and kept consistent for all replicates. Approximately 5 μL of sample was loaded into a glass capillary, which was then mounted on the XYZ-stage. Electric potential was delivered to the sample by a platinum wire inserted into the capillary and placed in contact with the solution. The voltage applied to the tip holder ranged from 0.60 to 1.2 kV for native samples, and 0.80 to 1.40 kV for denatured samples. Nine replicates in each ion mode were collected for all proteins with measurements spread across 8 days over 3 weeks. These replicates were conducted in pairs by collecting data in both ion modes consecutively. To accomplish this, a CIU experiment was collected in +ESI before the instrument polarity was changed to −ESI. A second CIU experiment was performed, beginning with −ESI to counteract any changes in protein stability due to pH changes during ESI. (14) This allowed for use of the same capillary, at the same position in relation to the instrument inlet, for each +ESI/–ESI replicate pair. A new tip was used for each +ESI/–ESI replicate pair. CIU experiments were accomplished by increasing collision voltage (CV) to the trap cell in 5 V steps, from 0 V to a maximum of 120 V.
Results and Discussion
Ten proteins were chosen for analysis in +ESI and −ESI in this work (Table 1). These proteins can be broken into three groups depending on the net-charge in solution at pH 7: net-negative (with pI < 6.5), net-neutral (with 6.5 ≤ pI ≤ 7.5), and net-positive (with pI > 7.5). Five of the ten proteins have a net-negative charge at pH 7, though four of these proteins (con A, β-lactoglobulin, ovalbumin, and α-lactalbumin) are routinely analyzed via +ESI-IMS-MS. (18,28) Additionally, each protein had charge-state distributions in +ESI and −ESI that resulted in at least one overlapping charge state (Figure 1), allowing for direct comparison of TWCCSN2→He and CIU50 in both ion modes.
 Anal. Chem. 2026, 98, 6, 4518–4527: Figure 1. Representative mass spectra in +ESI (top) and −ESI (bottom) for (A) ovalbumin, (B) GNA, (C) β-lactoglobulin, (D) α-lactalbumin, (E) con A, (F) streptavidin, (G) ubiquitin, (H) WGA, (I) cytochrome c, and (J) lysozyme. Proteins that have blue or red charge states indicate additional oligomeric states that were detected. Outlines in red boxes are overlapping charge states that are compared herein. *Indicates contaminant peaks.
Anal. Chem. 2026, 98, 6, 4518–4527: Figure 1. Representative mass spectra in +ESI (top) and −ESI (bottom) for (A) ovalbumin, (B) GNA, (C) β-lactoglobulin, (D) α-lactalbumin, (E) con A, (F) streptavidin, (G) ubiquitin, (H) WGA, (I) cytochrome c, and (J) lysozyme. Proteins that have blue or red charge states indicate additional oligomeric states that were detected. Outlines in red boxes are overlapping charge states that are compared herein. *Indicates contaminant peaks.
Proteins with Disulfide Bridges are Stable in Both Polarities
For both lysozyme and α-lactalbumin, the average TWCCSN2→He were within experimental uncertainty in +ESI and −ESI for both the initial, folded structures and the final, extended structures (Table 2). However, the 6+ and 6- ions of lysozyme and α-lactalbumin showed a high resistance to unfolding with >90% of the ions retaining their folded TWCCSN2→He values even at high CV (Figure 2). Replicate data showed similar trends (Figure S2). While CCS values provide a description of gas-phase structures, CIU experiments provide information on the stability of the gas-phase protein structures. Furthermore, the number and pattern of disulfide bridges present in antibodies has been characterized by CIU. (63) Lysozyme and α-lactalbumin both contain the glycosyl hydrolases family 22 (GH22) domain, which has four disulfide bridges that span ∼94% and ∼93% of the amino acid sequence, respectively (Figure S3). Thus, we hypothesize that rather than effects from protein pI and ionization polarity, the unfolding behavior of these proteins is controlled to a greater extent by the presence of stabilizing disulfide bridges that prevent routine unfolding during collisional activation.
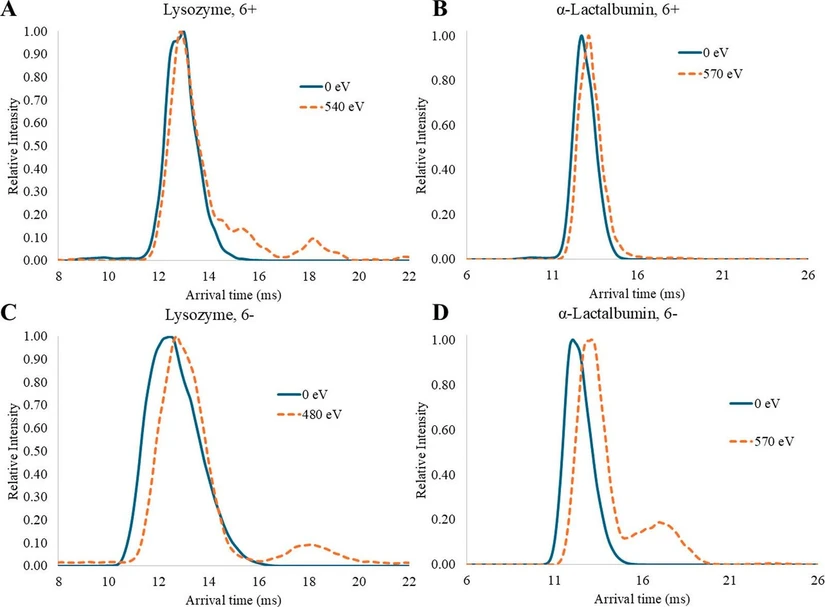 Anal. Chem. 2026, 98, 6, 4518–4527 - Figure 2. Representative mobiligrams showing arrival times for lysozyme (A, C) and α-lactalbumin (B, D) at low and high CV (blue solid and orange dashed lines, respectively) for +ESI (A, B) and −ESI (C, D). For lysozyme, different values were chosen for the high CV (e.g., 540 eV in +ESI and 480 eV in −ESI) because these were the last analyzed CV where the signal-to-noise ratio (S/N) for the 6 ± ions was greater than 3.
Anal. Chem. 2026, 98, 6, 4518–4527 - Figure 2. Representative mobiligrams showing arrival times for lysozyme (A, C) and α-lactalbumin (B, D) at low and high CV (blue solid and orange dashed lines, respectively) for +ESI (A, B) and −ESI (C, D). For lysozyme, different values were chosen for the high CV (e.g., 540 eV in +ESI and 480 eV in −ESI) because these were the last analyzed CV where the signal-to-noise ratio (S/N) for the 6 ± ions was greater than 3.
To characterize the stabilizing effect of disulfides, MD simulations were conducted that heat denatured 6+ ions of lysozyme and α-lactalbumin with disulfide bridges either present or removed with the resulting cysteine residues being protonated prior to heating (Figure S4). Structural changes associated with heat denaturation were characterized by taking the root-mean-square deviation (RMSD) of the heated structures relative to initial, crystal structures. Over ten trials, lysozyme ions possessed an average RMSD of (1.6 ± 0.2) nm when disulfides were present which increased to (3.5 ± 0.5) nm when disulfides were removed. Similarly, the RMSD of α-lactalbumin ions increased from (1.8 ± 0.1) nm when disulfides were present to (3.7 ± 0.3) nm when disulfides were removed, suggesting that the disulfides serve as a barrier to unfolding. This is visually represented in Figure S4C,D where simulated structures of heated lysozyme and α-lactalbumin ions with intact disulfides adopt more condensed configurations relative to those with disulfides removed. In essence, the disulfides “tie” disparate regions of protein together, which serves as a barrier to more thorough unfolding even at a maximum simulated temperature of 900 K, explaining the similarity of TWCCSN2→He values in +ESI versus −ESI as well as the limited percentage of ions that undergo a change in TWCCSN2→He upon additional collisional activation.
WGA has 16 disulfide bridges per monomeric subunit (Table 1). On average, these disulfides encompass seven amino acids each, with ∼78% the total amino acids in WGA being encompassed by at least one disulfide bridge (Figure S3). Compared to lysozyme and α-lactalbumin, the presence of disulfide bridges across small numbers of amino acids enables WGA to unfold with ΔCCS of (370 ± 60) Å2 and (390 ± 120) Å2 in +ESI and −ESI, respectively (Table 2). However, in comparison to the other lectins, GNA and con A, we hypothesize that these disulfide bridges in WGA limit the extent of unfolding, resulting in the extended, TWCCSN2→He values being within experimental uncertainty in both ion modes.
To summarize, lysozyme and α-lactalbumin exhibit similar gas-phase behavior in both +ESI and −ESI, with most ions retaining folded structures even under high CV. We hypothesize this resistance to unfolding is largely attributed to their four disulfide bridges, which span over 90% of their sequences and act as stabilizing constraints. MD simulations confirm that removing these disulfides significantly increases structural deviation during heat denaturation, indicating their critical role in maintaining compact configurations. Consequently, the presence of disulfide bridges explains the minimal changes in TWCCSN2→He and the proteins’ high stability under activation.
Conclusions
IMS-MS allows for detailed analysis of the structure and stability of native-like proteins in the gas phase. The structure of a protein is dictated by its amino acid sequence and the resulting noncovalent interactions that form the secondary, tertiary, and quaternary structures. Chargeable amino acid residues that are solvent exposed can carry charges leading to the protein having a net-positive, net-negative, or net-neutral charge in solution at physiological pH. Yet, most native IMS-MS analysis is conducted in +ESI, leading to proteins that are net-negative in solution experiencing large swings in their net-charge during the ESI process.
This work highlights that gas-phase stability of native-like proteins varies with ionization polarity. While CCS values for folded structures are generally comparable between +ESI and −ESI, unfolding energetics differ─net-negative proteins are more stable in −ESI, whereas net-positive proteins are more stable in +ESI. We hypothesize that these differences are due to changes in the intramolecular interactions within cationic and anionic proteins. Therefore, proteins analyzed in the mode opposite to their solution-phase charge may have different gas-phase structures compared to biological systems. For example, proteins that are net-negatively charged in physiological environments (pI < 6.5) are more likely to represent their biological counterparts as gas-phase anions when analyzed in −ESI compared to +ESI, enabling nMS experiments to more closely characterize the structures present in biological systems.